It is well known that when there are too many variables and a small sample size, many supervised classification algorithms tend to overfit the data. That is, even there is no actual difference between the groups, the program will still be able to discriminate them by picking some features that are “different” between these two group by pure chance! Of course, the classifier will be useless for new data since the pattern it detected is not real (not significant).
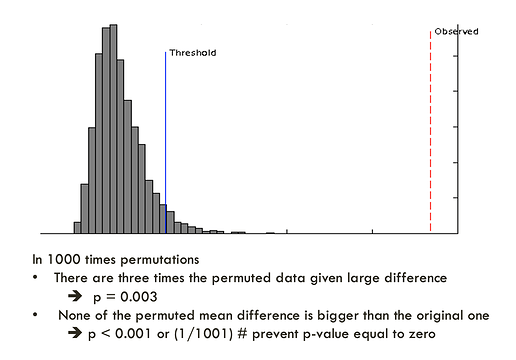
The purpose of a permutation test is to answer the question - “what is the performance if the groups are formed randomly”. The program uses the same data set with its class labels reassigned randomly. It then builds a new classifer, its performance is then evaluated. The process will repeat many times to estimate the distribution of the performance measure (which not necessarily follows a normal distribution). By comparing the performance using the original label and the performance based on the randomly labeled data, one can see if the former is significantly different from the latter. For PLS-DA, the performance is measured using prediction accuracy or group separation distance using the “B/W ratio” (as suggested by Bijlsma et al.). The further away to the right of the distribution formed by randomly permuted data, the more significant the discrimination. The p-value is calculated as the proportion of the times that class separation based on randomly labeled sample is at least as good as the one based on the original data (one-sided p value). An example output is shown below:
Related post: permutation tests in Biomarker Analysis