Hi,
Thank you for making such a fantastic tool to analyze LC-MS data. I have used Metaboanalyst successfully for a few years now, and have analyzed my specific datasets successfully numerous times.
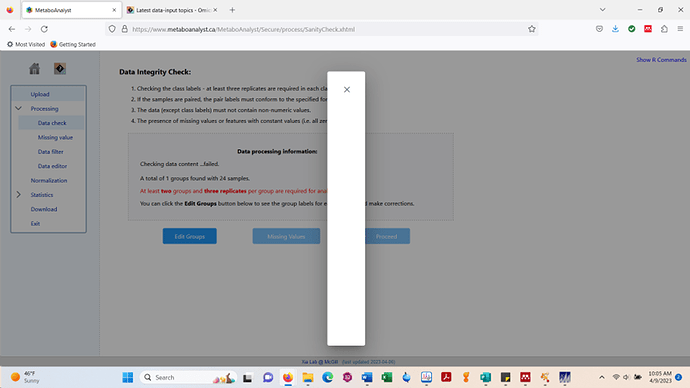
The last time I analyzed this specific data was in early March 2023. However, when I upload the same .csv today, I get a blank pop-up when I go to adjust my groups. I did not encounter this issue last month.
I have had intermittent problems like this occasionally, which has been resolved by using different browsers. However, I have this same issue whether I use Chrome, Firefox, or Edge. I also cleared my cache, just in case, to no avail.
Thanks for any insight,
Lydia
1 Like
I have got a problem like you
hope someone help
Thank you
Hi again,
Yes, the problem still persists this morning. Maybe the data uploader is the issue?
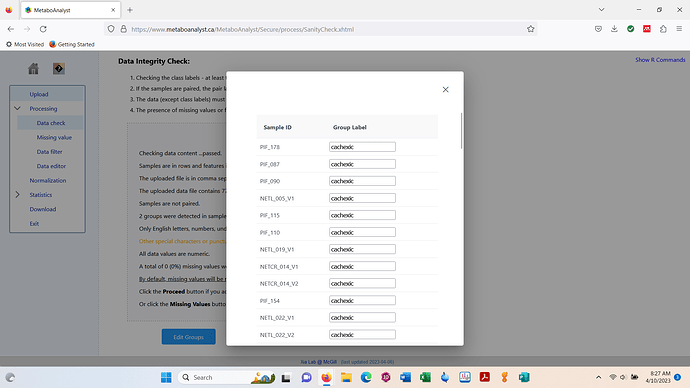
Test data looks fine. Screenshot below.
Thank you,
Lydia
Hi Lydia,
I have been having this same problem, have you been able to resolve it? I have it in every browser on multiple machines, but I have a colleague that uploaded the same .csv file and it was able to be modified. Maybe it’s the security software?
1.) Tool: Metaboanalyst (statistical analysis, one factor); module: .csv file uploader
2.) Reference .csv file is attached.
20230307_2022Met_feature_sub_dup_meladded_10885.csv (53.6 KB)
3.) Steps involved with issue:
a.) Navigate to the Metaboanalyst one-factor analysis upload page: MetaboAnalyst
b.) Adjust settings: ‘peak intensity’ > samples in columns (unpaired) > upload attached .csv > submit.
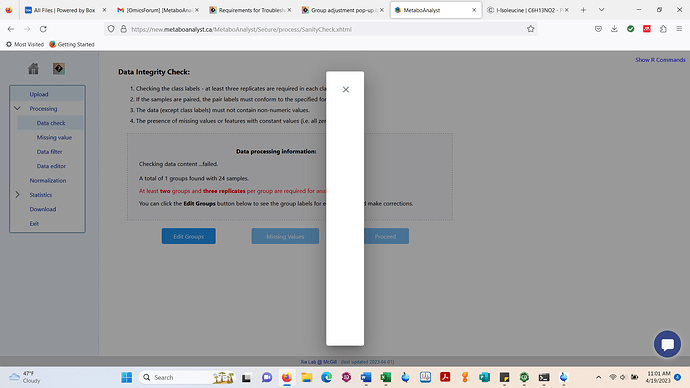
c.) Data integrity check page > edit groups > screenshot below is encountered on my end.
Thank you for your time and consideration.
Lydia
I don’t see the group information. The message (in red) actually tells the exact issue with your data. You can edit your labels to fix some minor issues - this is a major issue as the group label information is completely missing.
I edited my file to reflect the same labels as demonstrated in one of the sample datasets. Of course, this edited dataset could be analyzed without issue. I have attached both files for community purposes.
My edited dataset:
20230307_2022Met_feature_sub_dup_meladded_10885_test.csv (53.8 KB)
Based on sample dataset:
lcms_table.csv (48.4 KB)
Even though I have analyzed this specific data (unedited) without issue since last October, I just encountered this issue beginning in April 2023. Perhaps a new update made this a requirement.
Either way, thank you for your time and explanation, I hope this post will be helpful to peers.
Many thanks,
Lydia