Good afternoon,
I am having trouble with the compounds 2- and 3-phosphoglyceric acid. While they are recognized by the system, when looking into the pathway analysis, most specifically into glycolisis/glyconeogenesis pathways, they do not show up but rather the compound is presented in black as if they werent there.
Can you help me?
Thanks
Please refer to this post.
If this does not answer your question, please provide more details following our post guide
Hi again,
I have taken a look at the page you mentioned but I think that those compounds are included in the pathway libraries the error must be other ¿no? The compounds are recognized although they have no KEG associated:

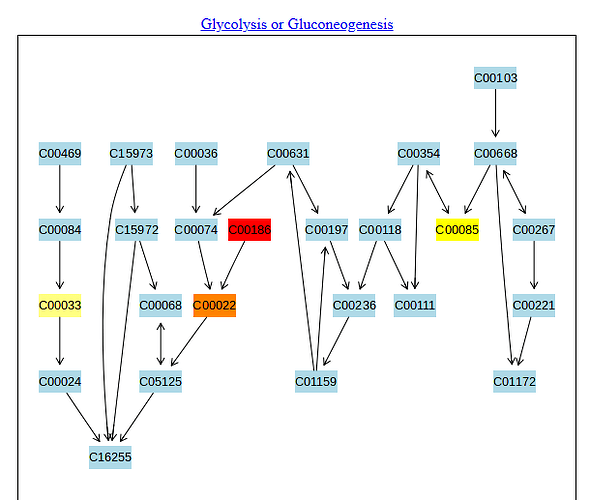
And then if you look into the pathway of glycolisis they remain as if the compound was not there (CD00631; 2-Phospho-D-Glycerate, and the same for CD00197, 3-Phospho-D-Glycerate) as you can see in the next image.
We have tried changing the name to the exact name that is shown in the pathway 2-phospho-D-glycerate, yet the same error happens, it is recognized as a metabolite but it is later not taken into accound on the pathway analysis, more specifically in glycolisis glyconeogenesis pathways.
Is it any way to solve this?