Yes, normalization and scaling are conducted before the fold-change and p-values are calculated, so they will impact these values. This is standard practice. You can see based on the order of the pages, and also you can click to see the R commands in the top right corner. This shows the order that different functions are called in.
However, if you are analyzing proteomics data, I suggest using ExpressAnalyst. It is a tool designed for gene expression analysis, also by our lab and using the same general design as MetaboAnalyst. We recently added a “proteomics” data input type, and have added proteomics-specific normalization methods. This way you can conduct pathway analysis on the results after, and also ExpressAnalyst accepts proteomics-specific IDs like Uniprot.
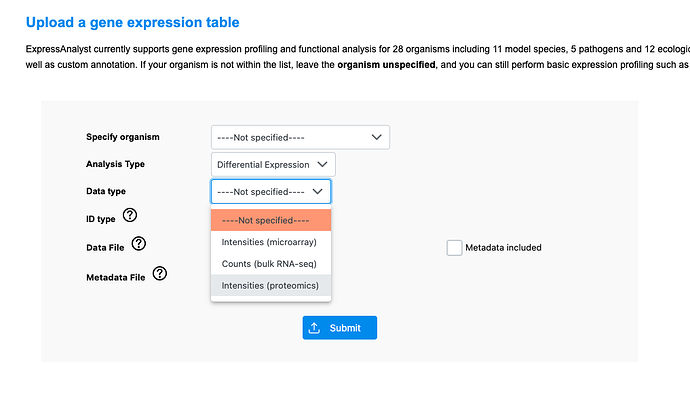
Go to the ‘single expression table’ module, then select the proteomics data input type: