Hi
I am Akanksha, working on Plant abiotic stress. Metaboanalyst is very user-friendly web-based tool. I have been using this tool from couple of weeks to interpret the dominant pathways associated with my Maize roots metabolomic study under drought (dehydration stress). For that I am simply pasting the list of compounds which are differential in my data. I have gone through the Omics forum to see the discussion related to my query and it was quite useful. I still have following questions:
-
For some compounds (eg. S-methylmethionine, pyroglutamine Sulphotyrosine, many lipids including 1-palmitoyl-2-linolenoyl-digalactosylglycerol (16:0/18:3), 1-linoleoyl-GPI (18:2), etc.) names are not matching and are excluded from the analysis even when I am trying to change names according to IUPAC nomenclature. Since I am trying to make the conclusion based on the enriched pathways obtained from Metaboanalyst-Pathway Analysis, I want to be sure if there is any way to include the missing metabolites as I feel missing metabolites may affect the impact of pathway associated with my data.
-
Is it fine to use Oryza sativa for Maize root study, how much impact does it make on pathway analysis to use metabolites extracted from Maize and pathway analyzed using Oryza sativa.
-
Details regarding the terms of Pathway table
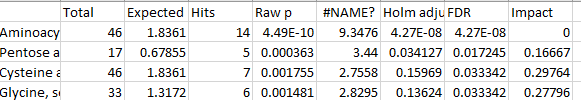
what does second column: Expected signifies?
difference between raw p value -log10 p value?
what value is Holm adjust? -
I am planning to include Overview of pathway Analysis figures in research article, and I am just making conclusion on the basis of obtained pathways according to list I provide (for example List of differential metabolites in common to case 1 and case 2, exclusive to case 1 and exclusive to case 2. Is it okay to do so or can you please suggest if there is any other way.
It would be really helpful if you could please suggest some explanation. Looking forward for your response.