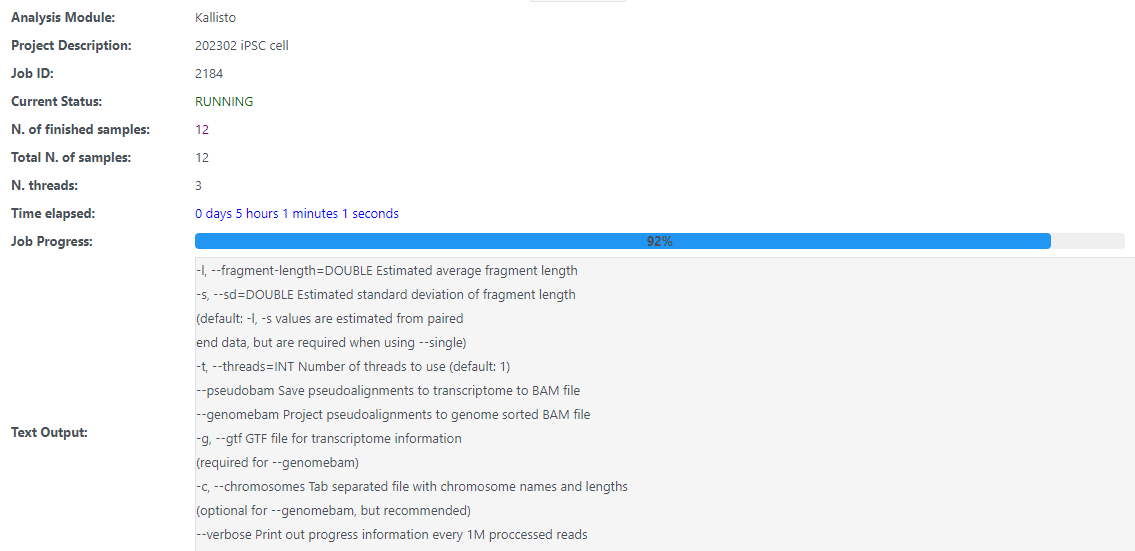
I have download the reference genome and put it in the database folder, but how to validate in kallisto?
Can you provide more details?
If you have a specific problem, please follow the post guidelines.
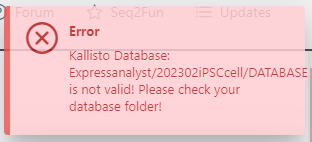
I got an error information after update the folders and files.
What reference genome did you download and what does the contents of your DATABASE folder look like?
The file is “hsapiens_GCF_000001405_NCBI.idx.gz” downloaded from NCBI, and there is no other files in the DATABASE folder.
Hi Jess, I worked out there is another folder in the DATABASE, and it is proceeding now. But it stuck on the Kallisto module for hours at 92% after finishing all the samples.
Which operating system are you using? We’ve been having some problems with Windows where Kallisto is running out of memory but not displaying errors through the interface.
Can you attach the job_logs.txt file from the RESULTS folder? It should have any error messages.
I found a error in the log, maybe I setup a wrong folder? Log is attached.
2023-02-17 13:34:53 Error in setwd(“/data/C:/User/li.sun/Expressanalyst/202302iPSCcell/RESULTS/results_2023_02_17_18_34_53”) :
2023-02-17 13:34:53 cannot change working directory
Log.txt (713.9 KB)
It looks like your file path is wrong somehow. See how it says “/data/C:/User …”? This is not a real filepath. Can you closely check the instructions on the DockerHub page, and make sure that you are: 1) using the right home directory parameter in the ‘docker run’ command and 2) putting your database/results folder in the right drive?
I also assume you are using Windows based on the “C:”.
This topic was automatically closed after 5 days. New replies are no longer allowed.