The parameter optimization in MetaboAnalyst is based on our OptiLCMS R package to achieve efficient parameter optimization using two main strategies - selecting high quality peaks for training and focusing on the most influential parameters. OptiLCMS includes three main steps:
-
Contaminant removal (optional)
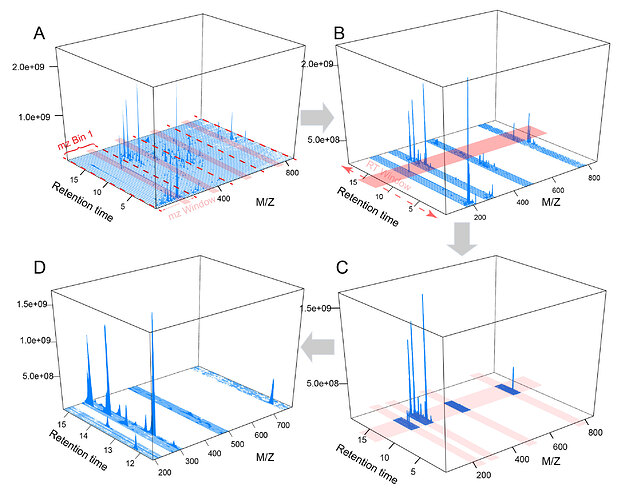
It is not rare that spectra data include some contaminants or noise from MS instruments or chromatography reagents. These mass signals usually appear persistently and may generate giant chromatographic peaks. These noise peaks should be excluded during the parameter optimization step. In OptiLCMS, all mass centroids are extracted and re-sorted from lowest to highest. All centroids that correspond to peaks that spread out over half of the whole chromatogram are excluded. Note, these centroids are only excluded during parameter optimization; they are not deleted from the raw spectra data. -
Regions of interest (ROI) extraction
Mass signals in LC-HRMS raw spectra are usually enriched in certain regions, rather than distributed evenly across the spectra. Parameter optimization based on the entire spectra is unnecessary. In OptiLCMS, a sliding window method is implemented in both m/z and retention time dimensions to extract multiple areas that are abundant with mass signals. These areas are the base for the subsequent parameter optimization stage. -
Parameter optimization
OptiLCMS focus on optimizing eight critical parameters used in the centWave algorithm. Parameters related to noise level (noise, prefilter value and prefilter abundance) and mass error (ppm) are estimated first using a kernel density estimator model. Then, a “Design of Experiment” model (central-composite model) is utilized to recursively estimate four other parameters (peak width, mzdiff, snthresh and bandwidth). Briefly, three levels (-1, 0 and 1) of all parameters are used to construct 44 combinations. The peak profiling results are evaluated based on the principal of selecting more stable and well-behaved peak groups. After the first round of optimization, a new round will be started by setting the best parameters from the last round as the initial values to optimize; this process is repeated until no better results can be obtained. The final optimized parameters will be used for peak profiling.
See more details from our previous publication.