Hi there, I’m having some issues using MetaboAnalyst 6.0 with the Statistical Analysis [metadata table] module, specific attempting to use the MEBA analysis in the Time-series only study design.
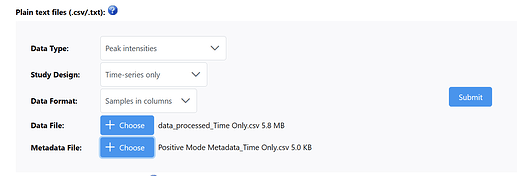
I’ve input my data type as Peak Intensities, study design as Time-series only, samples in columns and input my data and metadata files provided below:
Data File Link (it’s too big): data_processed_Time Only.csv
*If you go to file - export - export as csv on Onedrive you can download
Metadata File:
Positive Mode Metadata_Time Only.csv (5.0 KB)
The metadata processing passed, and I don’t edit anything on this screen and press proceed.
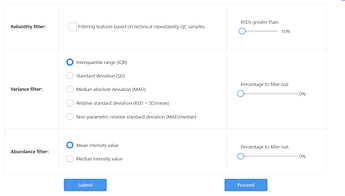
I do no data filtering as I’ve previously completed this, and press proceed without applying anything on Data Filtering as well.
For normalisation, I do normalisation by sum, log transformation and auto scaling.
All this works perfectly fine, and I’ve explored some of the other tests, but I’m interested in using the MEBA analysis. However, as soon as I click on the MEBA link it gives me Error: Please make sure data are balanced for time-series analysis. In particular, for each time point all experiments must exist and cannot be missing!
However, I’ve checked my data set several times and there are no missing values for any samples? It also showed 0% missing values at the data check. In this data set I have 200 samples, 5 time points (over 5 days so 5 repeats for each time point) and 8 subjects. I’m comparing 2553 features for their change over time, however I don’t think the large feature set is the issue, as I have an almost identical data set (same samples in negative ionisation) with 342 features and it still does has the error. I am a student very new to metabolomics analysis, any help is greatly appreciated! My apologies for the long post. I just wanted to make sure all details were included to be reproducible ![]()
Best,
Amanda