I would like to perform Enrichment and Pathway Analyses for my four experimental groups, but every time I try, I receive an error message. Is it possible to do these in MetaboAnalyst? Or is there any other tab that I could use?
Can you give us some more details? For example, a screen shot of what the error message looks like and more details on the format of your data.
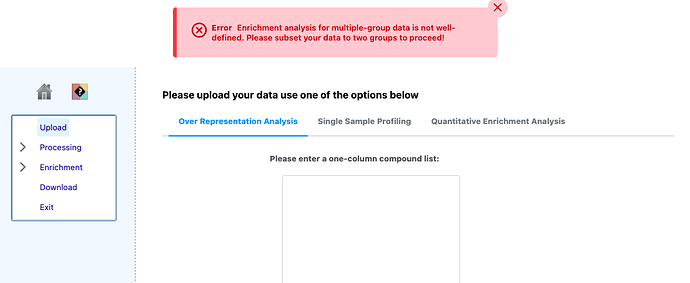
Sure. I am attaching the screen shot of the error message.
I have got 4 experimental groups (n is between 3 to 7 per group) with the area under curve values of the MS peaks for each sample. The sample IDs and groups are in the first two columns and metabolite or lipid names are on the rows. I have uploaded the data in the .csv format
Ok, this module is for one-column lists of compound names. You can click the example data to see what it should look like. You should have already done some statistical analysis to generate a list of metabolites of interest, and then you upload that here to see what pathways your list is enriched in.
If you have a different list for each experimental group, you can upload each list separately to get the enriched pathways for each.
If you still need to conduct statistical analysis (ie. identify metabolites of interest and then do enrichment analysis), you should start with the statistical analysis module in MetaboAnalyst and then upload your list here.
The error message is informative here. In general, enrichment analysis is only defined for two-group comparison - enrichment (or not) is always relative to a reference. When you give 4 groups, the program does not know which is the reference.
You can first perform ANOVA on four-group data (using Statistics Module), and then upload the significant metabolites (one column with one metabolite per row) for enrichment analysis here.