Dear all,
Some time ago I performed several analyses using MetaboAnalyst 5.0. We are currently running related experiments, so I wanted to check that I would obtain the same results using the new version of the software.
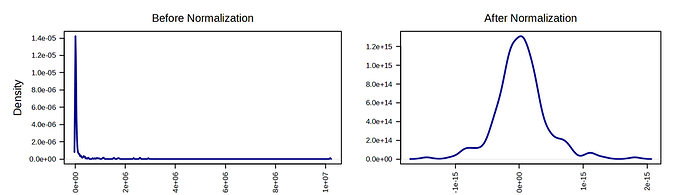
However, the analysis obtained with MetaboAnalyst 6.0 is different from the one previously obtained with version 5.0. As far as I can tell, I have followed exactly the same steps: log10 transformation and Pareto scaling.
I am not sure where the discrepancy might come from. I am attaching the analysis report generated previously with version 5.0 and the results obtained now with version 6.0, as well as the dataset used to generate the results.
I have also noticed that the commands shown at the end of the analysis history are slightly different in some aspects. However, as mentioned above, I have followed exactly the same workflow when performing the analysis. I should also mention that I am only using the online version of MetaboAnalyst, not the downloadable version.
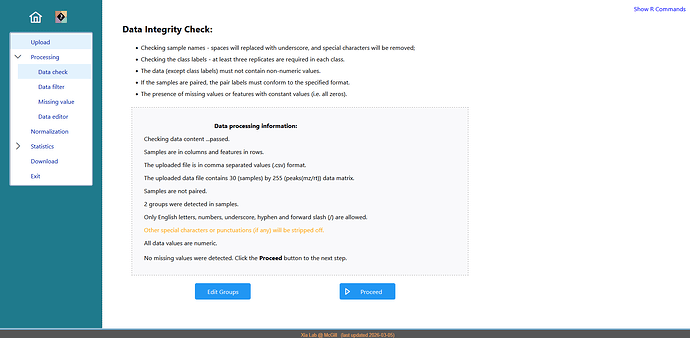
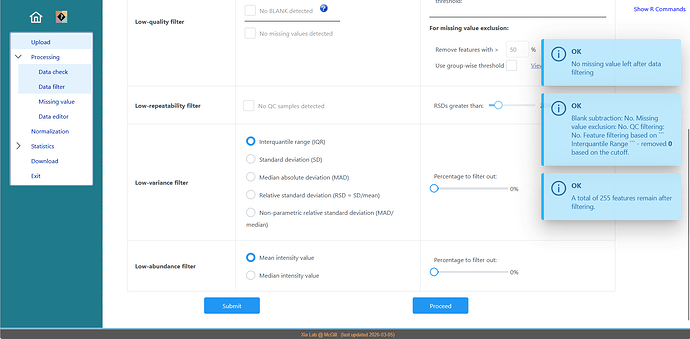
Additionally, in the previous analysis performed with version 5.0 no data filtering was applied. However, in version 6.0 the report indicates “Feature filtering based on Interquartile Range (IQR) – removed 0 features based on the cutoff”. However, I did not explicitly select any filtering step, so I am unsure why this appears in the workflow.
In addition, this dataset had never produced any errors before. However, when uploading it now I sometimes receive a message indicating that some metabolites cannot be read or that some entries are duplicated. The message does not appear every time, so the behaviour seems somewhat random. I have attached the message as well.
Could this difference be due to changes in the default preprocessing or filtering settings between versions 5.0 and 6.0?
Thank you very much in advance for your help.
Analysis_Report 5.0.pdf (711.5 KB)
Analysis_Report 6.0.pdf (6.8 MB)
dataset.csv (93.8 KB)