After removing singletons and constant features i have 361 ASVs remaining. I chose not to filter the data moving forward, as many of the (also rare) ASVs are eventually the same taxa. Altogether i have 93 different species.
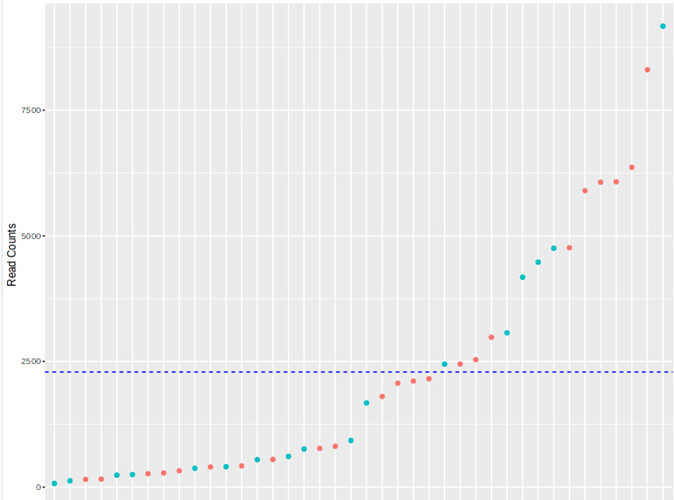
I also have very large differences in my lib sizes (see the image attached) with an average read count of 2295, lowest 72 and highes 9172
I am still struggling which normalisation method i should use for my data for diversity and differential abundance analysis, since i find no real consensus in the literature (concerning rarefaction, TSS, CSS,…). What is your recommendation for data normalisation?
Thank you for your help and support! I appreciate the work you do here!