Networks that contain too many nodes and edges are overwhelming, and difficult to use to draw clear biological conclusions. Thus, the feature selection step aims to consider only the most interesting features, and then to construct a network to understand the relationships between them. There are threeways of doing this: univariate differential analysis, multivariate dimension reduction and user-defined features.
- Univariate: use differential expressed features identified in “Data upload” page.
- Multivariate: use ranked list of feature contributions to each component from multivariate dimension reduction results.
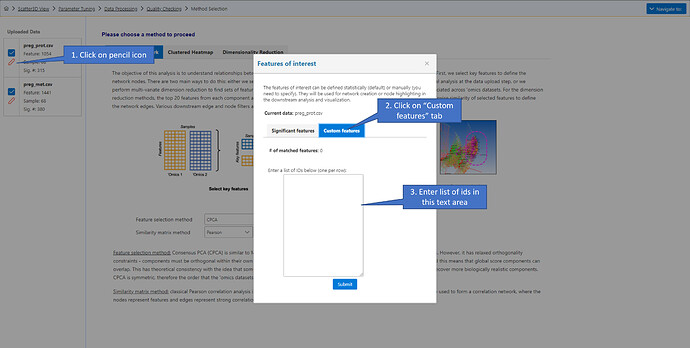
- User-defined: manually enter a list of features for each datasets, that will be used to query for correlation.