Hi, I’ve some challenges getting microbiome analyst to run the beta diversity. Thank you very much for taking a look at this issue.
- Tool and Module: MicrobiomeAnalyst - Marker Data Profiling
- I’ve attached a subset of the taxa and meta files. The attempt was saved under project ID 7990. Please contact me via email if the full data is required.
- Steps leading to problem
I’ve managed to upload the taxanomy included ASV table and its metadata, perform the data processing and all the usual steps including the standard low count and low variance filters, followed by CLR transformation.
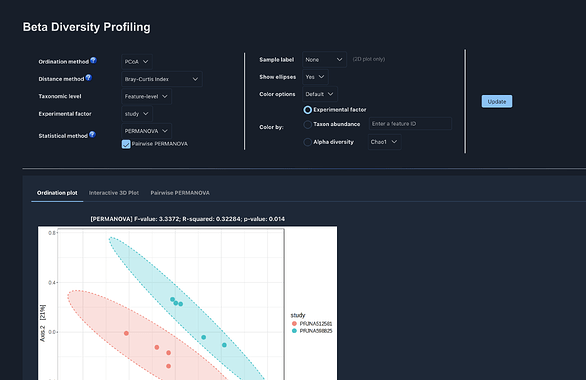
Other analyses such as Alpha diversity,random forest. its only the Beta diversity that is the issue
The log was:
[Success]: Login successful!
[Success]: Data has been uploaded sucessfully. All required files are present for analysis.
[Success]: A total of 292 low abundance features were removed based on prevalence.; A total of 9 low variance features were removed based on iqr; The number of features remains after the data filtering step: 76
[Success]: Normalization performed successfully!
[ERROR]:
Possible causes of error (last one being the most relevant):
[ERROR]: Could not find the image for downloading.
And the interface was:
Sample meta.csv (525 Bytes)
taxa sample.csv (70.6 KB)