Dear developers,
Thank you for developing and maintaining such an amazing tool.
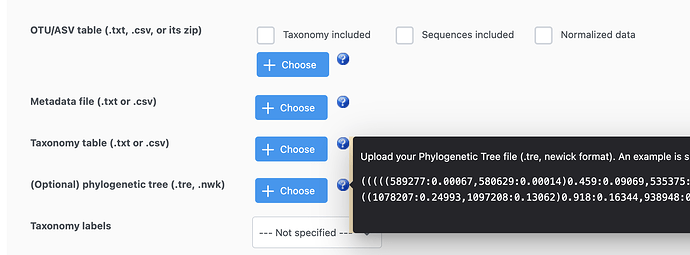
May I ask what will be the correct format of tree files for Unweighted/Weighted matrices in beta diversity analysis?
I tried the examples (with tree files) and they return the errors:
Tree tip labels do not match feature names in the OTU/taxonomic tables!
I am also attaching the tree files exported from QIIME2 with the scripts:
qiime phylogeny align-to-tree-mafft-fasttree
–i-sequences ./REPsequences.qza
–o-alignment ./treeFiles/aligned_seqs.qza
–o-masked-alignment ./treeFiles/aligned_masked.qza
–o-tree ./treeFiles/Unrootedtree.qza
–o-rooted-tree ./treeFiles/Rootedtree.qza
Then the rooted and unrooted tree files were exported with:
qiime tools export --input-path ./tree/Rootedtree.qza --output-path ./tree
Rootedtree.txt (2.5 MB)
Unrootedtree.txt (2.7 MB)
For uploading, I change the file extension from nwk to txt.
Best,
Yuping