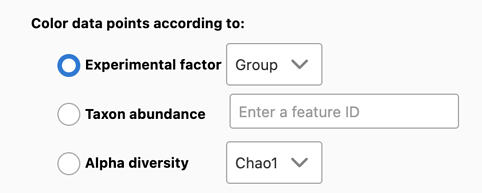
To facilitate pattern discovery, MicrobiomeAnalyst provides three options (see the Fig. below):
- Particular experimental factor from the meta-data (default option);
- Their alpha diversity measures (shown as color gradients);
- Abundance levels (shown as color gradients) of a particular feature (i.e. particular phylum, family, species, etc.) that they contain.
Note the use of this coloring option can be combined with the “label by” option to reveal more details about the available patterns.