Ranking the list of genes to be analyzed by GSEA is a critical step that can greatly influence the result. Many ranking metrics are present in the literature and there is no consensus on which is best to use in different scenarios. ExpressAnalyst offers four different methods.
- P-values: calculated based on the DE method used
- Fold changes (FC) FC with respect to the primary metadata factor;
- Moderated Welch’s t-test (MWT): MWT is a version of the t-test that allows for unequal variance;
- Signal-to-noise ratio (S2N): S2N is the difference between the mean expression divided by the sum of the expression standard deviation for two phenotype groups.
Please note the above gene ranking methods are not applicable to meta-analysis. Instead, the genes are ranked based on the summary statistic obtained from the previous meta-analysis (combine p-value, effect-size or direct merging). Results obtained from vote count can not be used to perform GSEA.
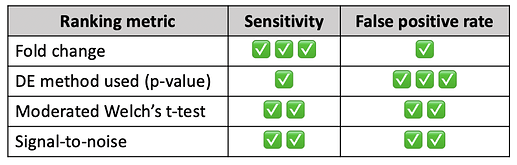
A recent publication compared different ranking metrics using 28 benchmark datasets and scored each one based on their sensitivity and false positive rate, summarized in the table below for the four metrics supported by ExpressAnalyst. While all metrics are widely accepted, you should choose based on how important the sensitivity/false positive rate is to your analysis.