LDA Effective Size (LEfSe) is a biomarker discovery and explanation method for high-dimensional data originally developed for microbiome data analysis. It integrates statistical significance with biological consistency (effect size) estimation.
- Non-parametric factorial Kruskal-Wallis (KW) sum-rank test to detect features with significant differential abundance with respect to the class of interest;
- Linear Discriminant Analysis to estimate the effect size of each differentially abundant features.
The result consists of a table listing all the features, the logarithmic value of the highest mean among all the classes, and if the feature is discriminative, the class with the highest mean and the logarithmic LDA score (Effect Size).
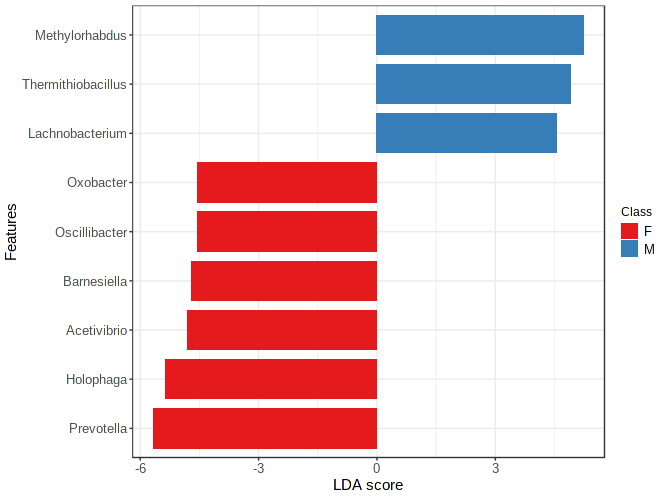
Please refer to the original paper by Segata et. al for more detailed explanations. Note there are some differences with regard to default parameter settings in the MicrobiomeAnalyst implementation of LEfSe. Please refer to this post.