Hello!
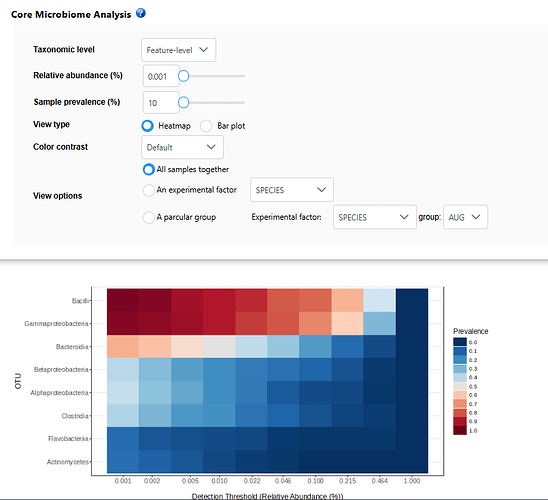
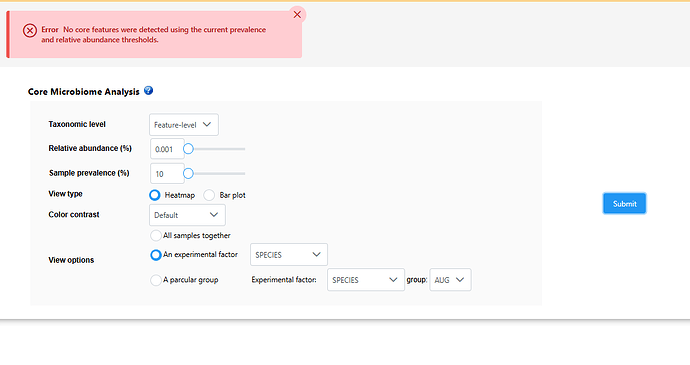
I keep getting error messages when attempting to get the core microbiome for experimental factor levels in my data. I can get a heatmap for all samples, though when I change the options to view for an experimental factor it comes up with error message “No core features were detected using the current prevalence and relative abundance thresholds”. I have reduced the prevalence and relative abundance to 0.001 and 10 respectively and still getting the error.
I have used the same data input to generate these figures before but cannot get them work again. I also tried to replicate this analysis using some of the provided data “mammalian gut” but am getting the same error message.
Thanks in advance ![]()
FLY_SAMPLES 1.txt (3.3 KB)
FLY_CLASS_TAXA 1.txt (14.4 KB)