Yes.
- Select the SNPs/metabolites of interest using the checkboxes in the node table on the left panel;
- Locate the “Highlight shared/all interactions” function on the table tool bar; choose Shared instead of the default “All” option;
- Click “Submit” button.
The shared nodes, the source nodes as well as their connections will be highlighted on the network. You can then perform enrichment analysis on these shared nodes.
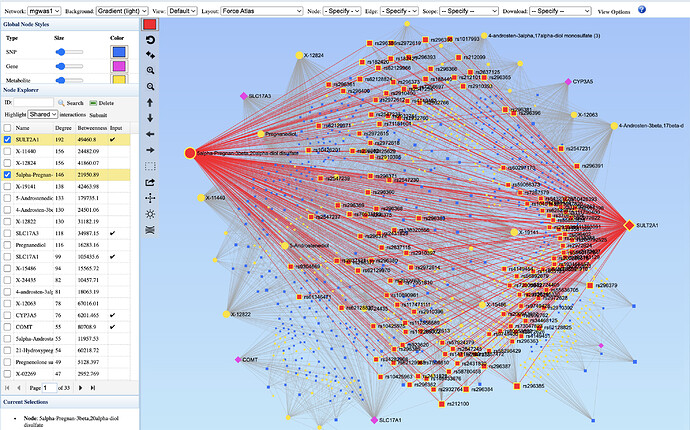
The screenshot below shows the shared SNPs between two nodes - SULT2A1 and 5alpha-Pregnan-3beta,20alpha-diol disulfate